The Effects of Obesity on Skeletal Muscle Contractile Function

Skeletal muscle
Given that obesity has profound effects on all organ systems, it is not surprising that excess body fat is also associated with a decrease in muscle function.
The complex biology of the molecular, structural, and functional changes that have been associated with obesity are now extensively discussed in a review article by James Tallin and colleagues, published in the Journal of Experimental Biology.
Without going into the molecular details here, suffice it to say that there is considerable evidence to show and explain why muscular function is impaired in both animal models and humans with excess body fat. (For e.g. at a cellular level, the dominant effects of obesity are disrupted calcium signalling and 5′-adenosine monophosphate-activated protein kinase (AMPK) activity. As a result, there is a shift from slow to fast muscle fibre types. There is also evidence for an impairment in myogenesis resulting from disruption of muscle satellite cell activation. Furthermore, muscle function is affected by insulin resistance and decreased adiponectin levels generally associated with obesity).
Although individuals with obesity will often have a larger muscle mass and may well be stronger than “normal-weight” individuals, when corrected for the amount of extra muscle, it is evident that the muscles are less efficient.
In fact, many of the biochemical and structural changes that occur in obesity are very similar to those found with aging. Not surprisingly, when aging meets obesity, things get even worse.
Although the paper does not discuss the reversibility of these changes with weight loss (or obesity treatment in general), I am aware of other data showing that much of the loss of muscle contractile function associated with obesity can be reversed with weight loss.
A clinical correlate of this is the fact that, following weight loss, individuals often find that it takes far more exercise to burn the same number of calories than before (this is not just because the person is now carrying less weight).
Given the increased recognition that lean body mass is an important determinant of overall health and function, clearly this topic is of continuing interest.
@DrSharma
Edmonton, AB
Factors Affecting Energy Metabolism: Genetics, Sex, and Aging
 Continuing with citations from my article in Obesity Reviews on an aeteological framework for assessing obesity, we now turn to the some of the factors that can affect metabolic rate:
Continuing with citations from my article in Obesity Reviews on an aeteological framework for assessing obesity, we now turn to the some of the factors that can affect metabolic rate:
Genetics
Because heritable factors appear to be responsible for 45–75% of the inter‐individual variation in body mass index (BMI), the potential impact of genetic determinants of metabolic rate upon the predisposition to obesity must be considered. While numerous somatic and mitochondrial genes with potential effects on metabolic rate have been identified, their contribution to human obesity has yet to be defined Likewise, although there is preliminary evidence for intrauterine and perinatal programming of genes involved in energy metabolism, their role in human obesity remain unclear. What is apparent is that the genetic predisposition to obesity (including both energy intake and metabolism) is not explainable on the basis of a small number of common mutations exerting substantial effects on the individual tendency to weight gain. Thus, a great deal of work is still required before investigation into the multitude of genetic determinants of body weight can potentially impact clinical management. Currently, a careful clinical assessment of family history of obesity and related risk factors remains the best measure of genetic risk for obesity.
Sex
There is a clear effect of gender [sic] on metabolic requirements, whereby, for the same BMI, women consistently display lower metabolic rates (approximately 20% less) than men, largely accounted for by differences in fat‐free mass (FFM).
Aging
Aging is an important determinant of a decline in metabolic rate, with an estimated reduction of around 150 kcal per decade of adult life. Factors that result in the age‐related decline in energy requirements include changes in neuroendocrine factors (e.g. sympathetic activity, thyroid function, etc.) as well as a reduction in skeletal muscle quantity and quality (resulting from reduced physical activity, reduced protein intake and other less‐well‐understood factors).
Additional factors that can affect metabolic rate will be discussed in subsequent posts.
@DrSharma
Edmonton, AB
The Heterogeneity of Obesity
 In the same manner in that there is not one predisposing factor for the development of obesity, the phenotypic clinical presentation of obesity is likewise extraordinarily heterogenous. (This has some authors speaking of “obesities” rather than “obesity”).
In the same manner in that there is not one predisposing factor for the development of obesity, the phenotypic clinical presentation of obesity is likewise extraordinarily heterogenous. (This has some authors speaking of “obesities” rather than “obesity”).
While it is now well established that BMI is a measure of size rather than health, it is perhaps less well recognised how the different types of body fat and their storage in various fat depots and organs can contribute to cardiometabolic disease (location, location, location!).
Now, a comprehensive review by Ian Neeland from the University of Texas Southwestern Medical Center, Dallas, together with my colleagues Paul Poirier and JP Despres from Laval University in Quebec, published in Circulation discusses the cardiovascular and metabolic heterogeneity of obesity.
As the authors point out,
“Although the BMI has been a convenient and simple index to monitor the growth in obesity prevalence at the population level, many metabolic and clinical studies have revealed that obesity, when defined on the basis of the BMI alone, is a remarkably heterogeneous condition. For instance, patients with similar body weight or BMI values have been shown to display markedly different comorbidities and levels of health risk.”
Not only has BMI never emerged as a significant component in risk engines such as the Framingham risk score, there are many individuals with obesity who never develop metabolic complications or heart disease during the course of their life.
The paper offers a good review of what the author describe as adipose dysfunction or “adiposopathy” = “sick fat”. Thus, in some individuals, there is an accumulation of “unhealthy” fat (particularly visceral and ectopic fat), whereas in others, excess fat predominantly consists of “healthy” fat (predominantly in subcutaneous depots such as the hips and thighs).
The authors thus emphasise the importance of measuring fat location with methods ranging from simple anthropometric measures (e.g. waist circumference) to comprehensive imaging techniques (e.g. MRI).
The authors also provide a succinct overview of exactly how this “sick fat” contributes to cardiometabolic risk and briefly touches on the behavioural, medical, and surgical management of patients with obesity and elevated cardiometabolic risk.
I, for one, was also happy to see the inclusion of the Edmonton Obesity Staging System in their reflections on this complex issue.
This paper is certainly suggested reading for anyone interested in the link between obesity and cardiovascular disease.
@DrSharma
Edmonton, AB
Can Liraglutide Help Grow New Fat Cells?
 The human GLP-1 analogue liraglutide is now approved for the long-term medical treatment of obesity in an ever-increasing number of countries. Its safety and clinical effectiveness is now well established and there is no doubt that this is an important addition to the rather limited number of treatment options available to people living with obesity.
The human GLP-1 analogue liraglutide is now approved for the long-term medical treatment of obesity in an ever-increasing number of countries. Its safety and clinical effectiveness is now well established and there is no doubt that this is an important addition to the rather limited number of treatment options available to people living with obesity.
Interestingly, however, liraglutide has also been shown to promote the differentiation of pre-adipocytes or, in other words, promote the formation of new fat cells.
While this may seen worrying or even counter-intuitive, we much remember that having more (smaller) rather than fewer (bigger) fat cells actually has substantial metabolic advantage s- there is indeed ample data showing that large adipocyte cell size and limited capacity to grow fat cells (the extreme case of which is seen in people with lipodystrophy) is actually a key risk factor for metabolic problems including insulin resistance, possible by promoting the accumulation of ectopic fat (e.g. in liver and skeletal muscle).
Now, a paper by Yongmei Liand colleagues, published in Molecular Medicine Reports provides additional insight into the cellular pathways involved in liraglutide’s adipogenic effects.
Using a series of in vitro experiments, the researchers show that liraglutide does indeed promote the adipogenic differentiation of 3T3-L1 cells (a widely used murine preadipocyte cell line) through a process that upregulates the expression of C/EBPα and PPARγ at the early phase of adipogenic differentiation, promots the expression of lipogenesis associated genes including aP2, and enhances the accumulated of lipids.
At the same time, liraglutide appears to suppress cell proliferation via the Hippo‑yes‑associated protein (YAP) signaling pathway, thereby allowing these cells to transform into mature adipocytes sooner.
How relevant these observations are for humans remains to be seen, but certainly the promotion of adipogenic differentiation may hold the potential for improving insulin sensitivity and reducing the metabolic risks associated with excess weight gain.
@DrSharma
Edmonton, AB
Disclaimer: I have received speaking and consulting honoraria from Novo Nordisk, the maker of liraglutide.
Molecular Changes During Weight Gain – Everyone Is Different
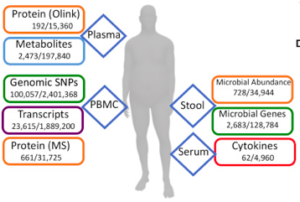 As one may well imagine, changes in body weight (up or down) can profoundly affect a vast number of hormonal and metabolic pathways.
As one may well imagine, changes in body weight (up or down) can profoundly affect a vast number of hormonal and metabolic pathways.
Now, a team of researchers led by Brian Piening and colleagues, in a paper published in Cell Systems used a broad “omics” based approach to study what happens when people lose ore gain weight.
Specifically, the goal of this study was to:
(1) assemble a comprehensive map of the molecular changes in humans (in circulating blood as well as the microbiome) that occur over the course of a carefully controlled weight gain and their reversibility with weight loss; and
(2) determine whether inulin sensitive (IS) and insulin resistant (IR) individuals who are matched for degree of obesity demonstrate unique biomolecular signatures and/or pathway activation during similar weight gain.
The study included 23 carefully selected healthy participants with BMI 25–35 kg/m2, were studied. Samples were collected at baseline. They then underwent a 30-day weight gain period (average 2.8 kg), followed by an eucaloric diet for 7 days, at which point a second fasted sample of blood and stool was collected. Each participant then underwent a caloric-restricted diet under nutritionist supervision for a subsequent 60-day period designed to return each participant back to his/her initial baseline weight, at which point a third set of fasted samples of blood and stool were collected. A subset of participants returned for a follow-up sampling approximately 3 months after the end of the perturbation.Insulin resistance was assessed at baseline using a modified insulin suppression test.
The large-scale multi-omics assays performed at all time points on each participant included genomics, proteomics, metabolomics and microbiomics.
Despite some differences between the IS and IR group (particularly in differential regulation of inflammatory/immune response pathways), overall, molecular changes were dominated by inter-personal variation (i.e. changes within the same individual), which accounted for more than 90% of the observed variance in some cases (e.g., cytokines). The most striking changes with weight gain were in inflammation response pathways (despite the rather modest weight gain) and were (fortunately) reversed by weight loss.
As the authors note,
“Comparing the variation in cytokine levels between multiple baselines in a single individual versus across individuals, we observed a striking difference: for almost all cytokines, the within-individual coefficient of variation was under 20%, whereas the variation across individuals was 40%–60%. This shows that our baseline cytokine profiles are unique to the individual, a point that has significant implications for one-size-fits-all clinical cytokine assays for the detection and/or monitoring of disease.”
On the opposite side of the spectrum, proteomics and metabolomics measurements had a substantial unexplained component (30% and 35%, respectively), highlighting the presence of unaccounted factors (e.g., food, exercise, and other changing environmental factors) or a subject-specific reaction to the perturbation.
Notably, not all of the responses we observed were consistent across IR and IS participants.
“In particular, for the microbiome, we observed that the microbe A. muciniphila was weight gain responsive only in insulin-sensitive participants. The abundance of this particular microbe in IR individuals did not change across perturbations and was barely or not detectable in most IR individuals.”
Clearly, these findings highlight the fact that each individual is biochemically unique, which the authors note, makes a strong case for personalized analysis in medicine.
Perhaps more importantly for researchers, nearly all of the data are publicly available, enabling exploration of inter-omic relationships and alterations across a longitudinal perturbation, thus providing a valuable resource for the development and validation of bioinformatic tools and pipelines integrating disparate data types.
@DrSharms
Edmonton, AB


