Transcriptional Control of Energy Regulation
Friday, October 30, 2015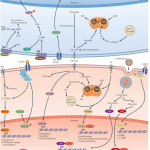 To students of human physiology, the commonly held view that obesity is simply a matter of energy in and energy out is nothing short of laughable.
To students of human physiology, the commonly held view that obesity is simply a matter of energy in and energy out is nothing short of laughable.
Indeed, there are perhaps no other biological functions of more importance for survival of an organism, than those that regulate energy uptake, storage and expenditure – functions, without any form of life would be impossible.
Thus, the finely tuned complex and often highly redundant pathways that have evolved to optimize energy metabolism have evolved to readily switch from states of feeding to starvation with shifts in substrate use (both qualitative and quantitative) – functions that are controlled by hundreds (if not thousands) of genes.
Getting these genes to work in concert, requires a complex system of gene regulation, by which individual genes are switched on an off (to allow or stop protein synthesis) in various tissues to just the right amount at just the right time – a process known as transcriptional control.
Now, a comprehensive review by Adelheid Lempradl and colleagues, published in Nature Genetics, summarizes the multitude of interlinked processes that control transcription of genes involved in energy homeostasis.
As the authors explain,
“Transcriptional control is the sum of the cellular events that select and dose gene transcription. In simple terms, these events converge on the regulation of gene locus accessibility and polymerase activity (including recruitment, pausing, processivity and termination).”
“Energy homeostasis requires multi-layered regulation via dynamic, often periodic, expression of metabolic pathways to properly anticipate and respond to shifts in energy state.”
“Transcription factors act by binding to specific regulatory DNA sequences, thus controlling the transcriptional output of defined target gene sets. They cooperate with co-regulators, which either promote (co-activators) or inhibit (co-repressors) transcription. Together, they build feedback networks and control the stability and responsiveness of energy homeostasis. Metabolic cells use receptors and metabolic machinery to generate specific signalling responses to endocrine inputs (for example, insulin, glucagon or leptin receptors) or metabolic inputs (for example, the primary energy metabolism machinery itself).”
The papers goes on to discuss at length the various regulator, co-regulators and the plethora of epigenetic modifiers that determine how these factors do their job of activating or deactivating relevant genes throughout the body.
Why is any of this important?
“Rapid progress is currently being made in research on chromatin-based regulation of gene expression. Particular unknowns include the mechanisms that establish long-term set points or priming of gene expression. Identifying the processes that establish activation thresholds and maximal output set points, as well as their self-organizing principles is currently an intriguing area of research, and is important in understanding susceptibility to complex trait disorders, including metabolic and autoimmune diseases. For example, many intergenerational and developmental reprogramming paradigms elicit metabolic disease susceptibility. They highlight the potential impact of subtly divergent transcriptional profiles in any given genetic context.”
In other words, understanding these processes are fundamental for our understanding of everything from the body’s weight set point (and how this is altered) to intergenerational transmission of obesity risk from one generation to the next (perhaps the most important driver of the current obesity epidemic).
But the complexity of these processes also raise important issues for clinicians,
“The seemingly exponential growth in this complexity now poses a major challenge for translational researchers in need of simplified but accurate paradigms for clinical use.”
The least we can do is to stop pretending that there is anything easy about energy in and energy out.
@DrSharma
Berlin, Germany


